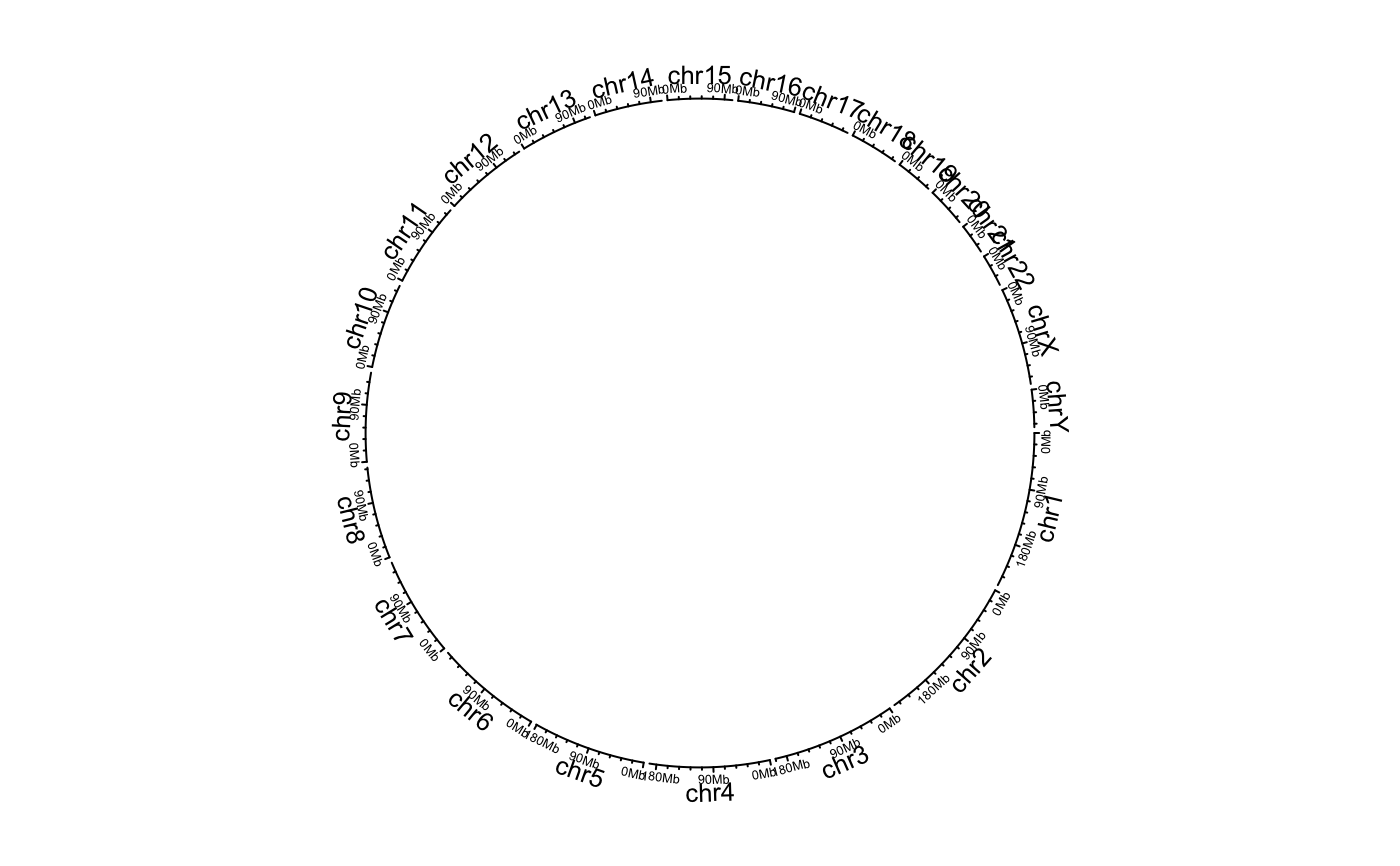
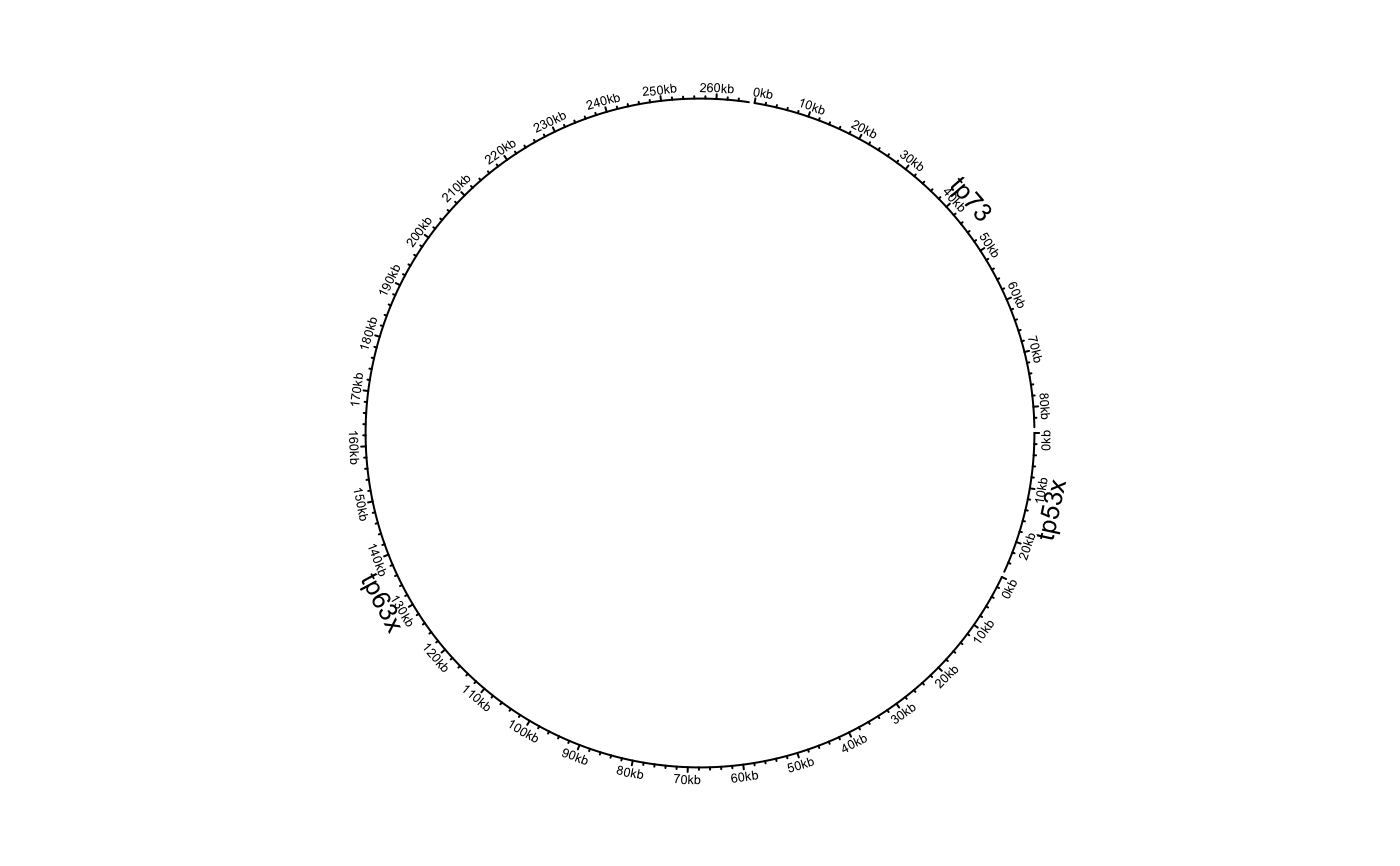
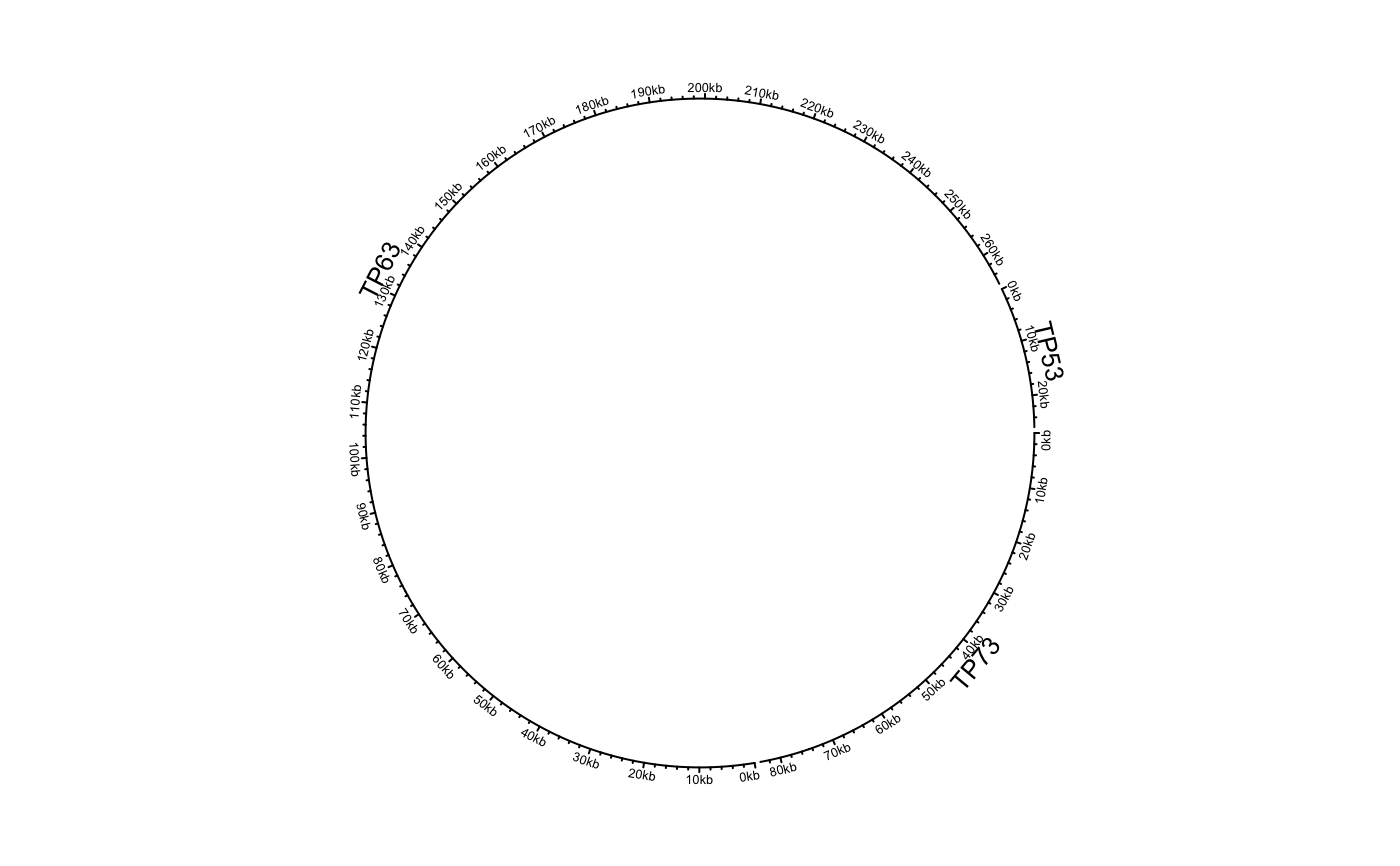
Initialize circular plot with any genomic data
circos.genomicInitialize.RdInitialize circular plot with any genomic data
circos.genomicInitialize( data, sector.names = NULL, major.by = NULL, plotType = c("axis", "labels"), tickLabelsStartFromZero = TRUE, axis.labels.cex = 0.4*par("cex"), labels.cex = 0.8*par("cex"), track.height = NULL, ...)
Arguments
| data | A data frame in bed format. |
|---|---|
| sector.names | Labels for each sectors which will be drawn along each sector. It will not modify values of sector index. |
| major.by | Increment of major ticks. It is calculated automatically if the value is not set (about every 10 degrees there is a major tick). |
| plotType | If it is not |
| tickLabelsStartFromZero | Whether axis tick labels start from 0? This will only affect the axis labels while not affect x-values in cells. |
| axis.labels.cex | The font size for the axis tick labels. |
| labels.cex | The font size for the labels. |
| track.height | If |
| ... | Pass to |
Details
The function will initialize circular plot from genomic data. If plotType is set with value in axis or labels, there will
create a new track.
The order of sectors related to data structure of data. If the first column in data is a factor, the order of sectors
is levels(data[[1]]); If the first column is just a simple vector, the order of sectors is unique(data[[1]].
For more details on initializing genomic plot, please refer to the vignettes.
See also
Examples
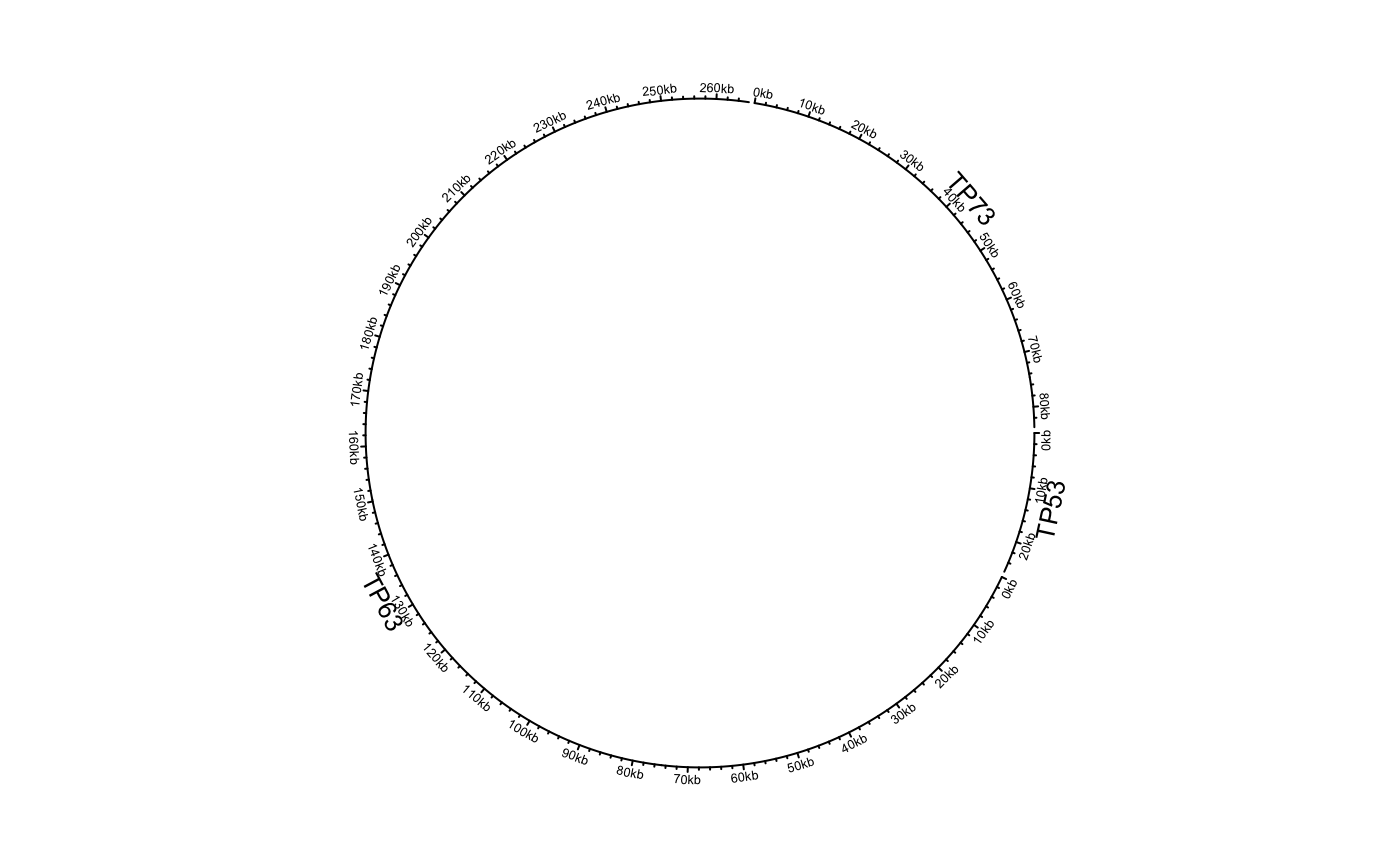
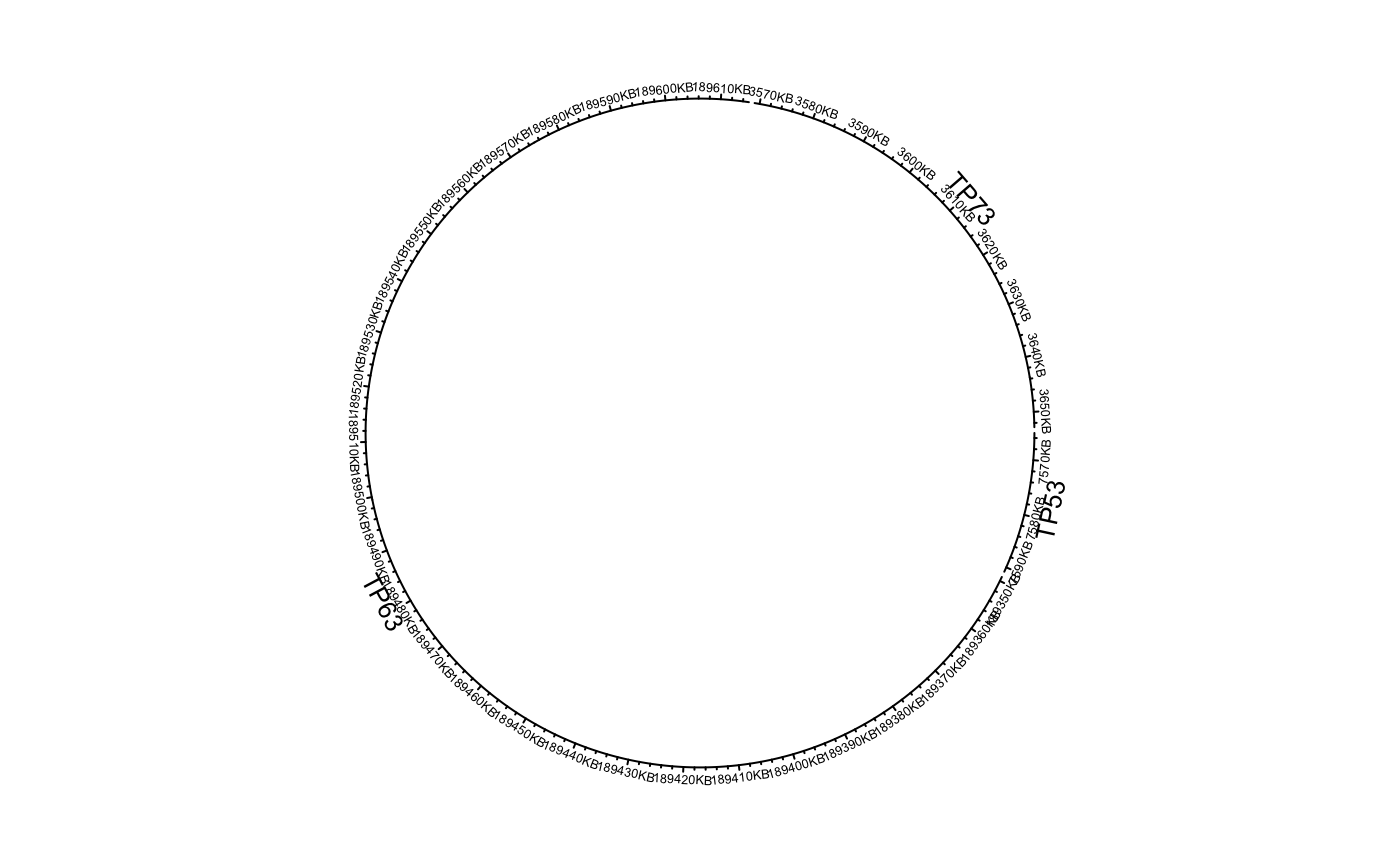
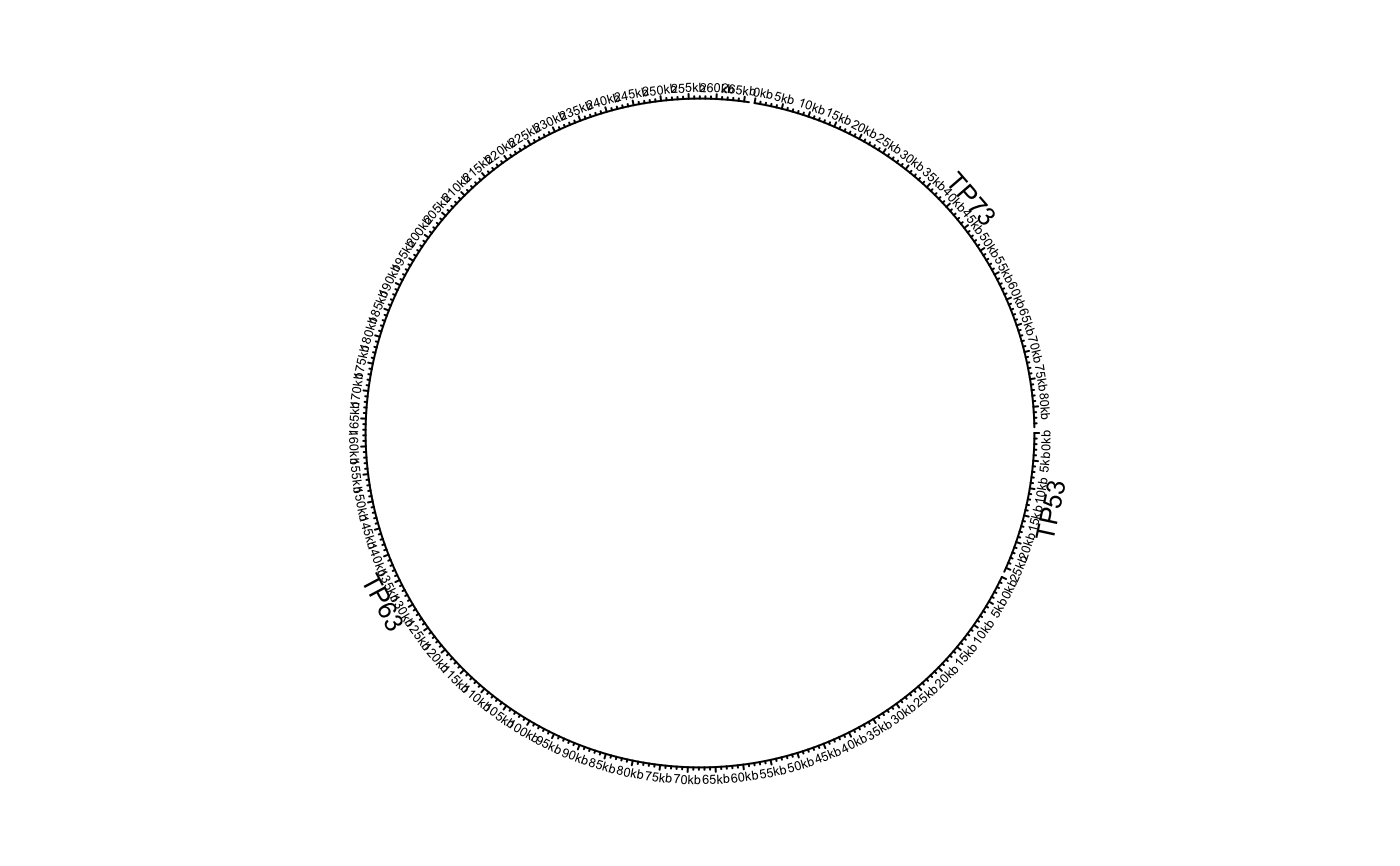
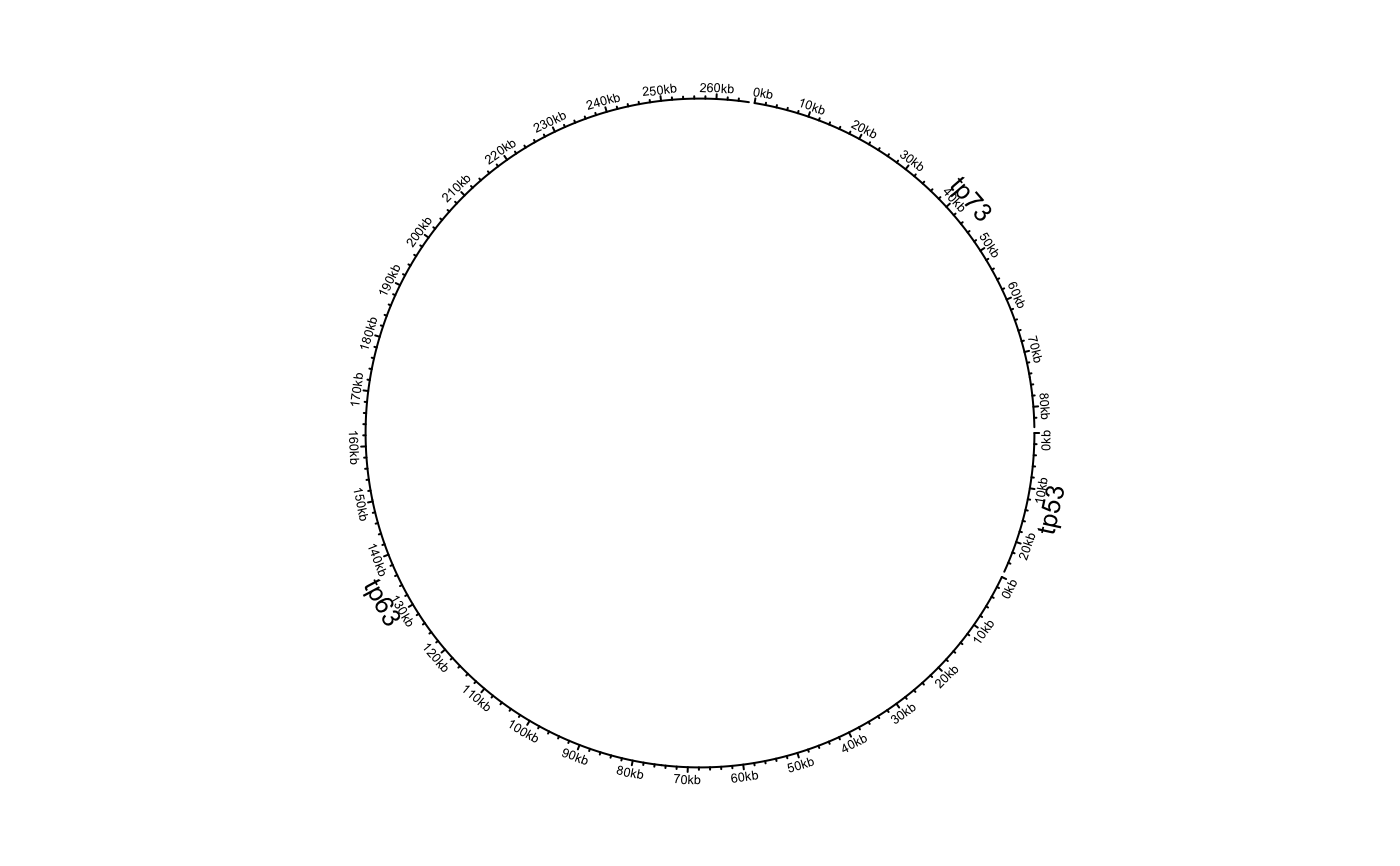
df = data.frame(name = c("TP53", "TP63", "TP73"), start = c(7565097, 189349205, 3569084), end = c(7590856, 189615068, 3652765), stringsAsFactors = FALSE) circos.genomicInitialize(df)circos.clear() df[[1]] = factor(df[[1]], levels = c("TP73", "TP63", "TP53")) circos.genomicInitialize(df)