Convert a phylo object to a dendrogram object
phylo_to_dendrogram.RdConvert a phylo object to a dendrogram object
phylo_to_dendrogram(obj, log = FALSE)Arguments
- obj
A
phyloobject.- log
Whether the height of the phylogenetic tree should be log-transformed (log10(x + 1)).
Details
The motivation is that phylogenetic tree may contain polytomies, which means at a certain node, there are more than two children branches. Available tools that do the conversion only support binary trees.
The returned dendrogram object is not in its standard format which means it can not be properly
drawn by the plot.dendrogram function. However, you can still apply dendextend::cutree to the returned
dendrogram object with no problem and the dendrogram can be properly drawn with the ComplexHeatmap package.
Value
A dendrogram object.
Examples
require(ape)
#> Loading required package: ape
data(bird.families)
d = phylo_to_dendrogram(bird.families)
require(ComplexHeatmap)
#> Loading required package: ComplexHeatmap
#> ========================================
#> ComplexHeatmap version 2.18.0
#> Bioconductor page: http://bioconductor.org/packages/ComplexHeatmap/
#> Github page: https://github.com/jokergoo/ComplexHeatmap
#> Documentation: http://jokergoo.github.io/ComplexHeatmap-reference
#>
#> If you use it in published research, please cite either one:
#> - Gu, Z. Complex Heatmap Visualization. iMeta 2022.
#> - Gu, Z. Complex heatmaps reveal patterns and correlations in multidimensional
#> genomic data. Bioinformatics 2016.
#>
#>
#> The new InteractiveComplexHeatmap package can directly export static
#> complex heatmaps into an interactive Shiny app with zero effort. Have a try!
#>
#> This message can be suppressed by:
#> suppressPackageStartupMessages(library(ComplexHeatmap))
#> ========================================
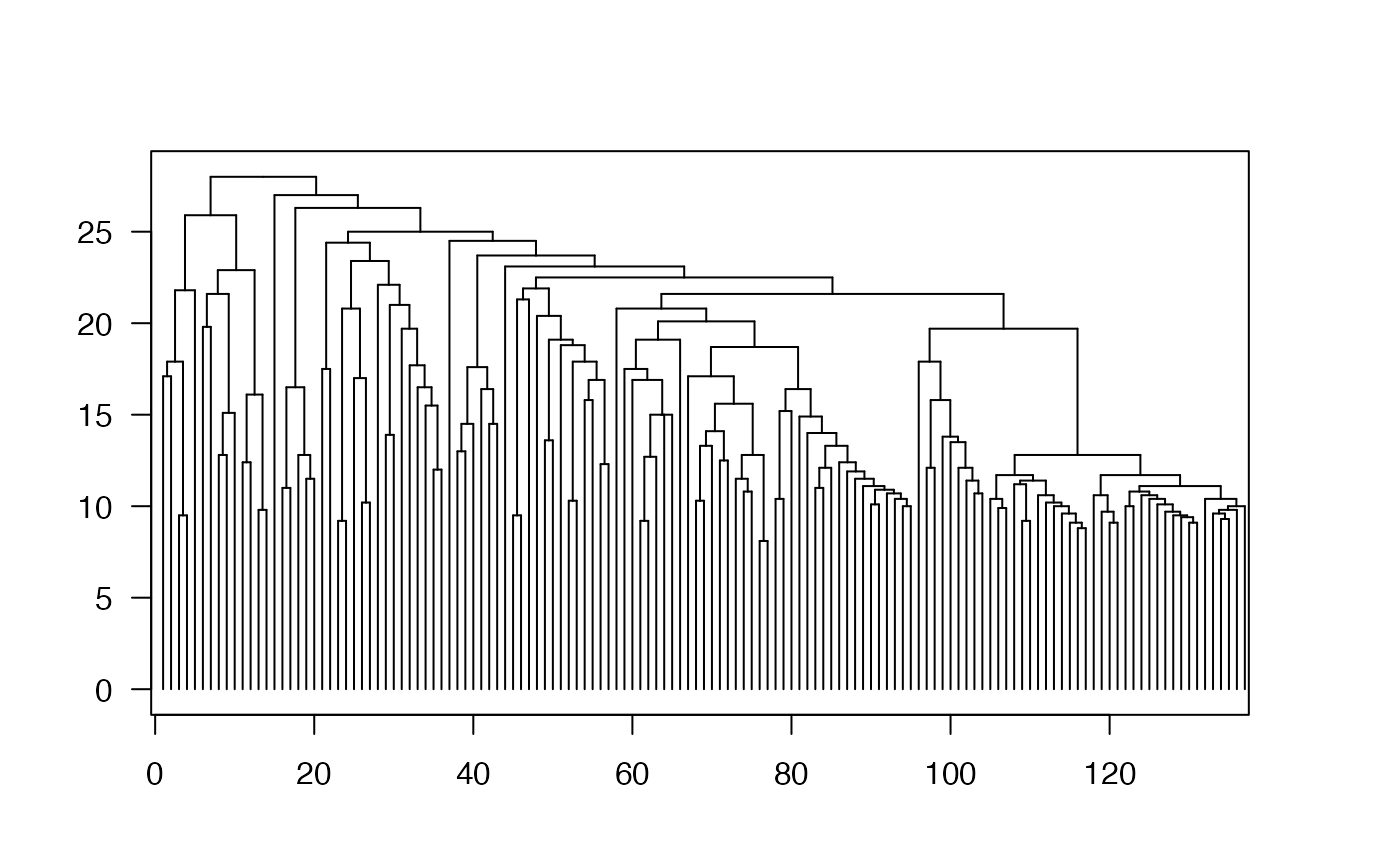
grid.dendrogram(d, test = TRUE)