ontology_DAG: a class for ontology data
Zuguang Gu ( z.gu@dkfz.de )
2024-09-13
Source:vignettes/v01_dag.Rmd
v01_dag.RmdOntologies are represented in a form of directed acyclic diagram (DAG). A DAG is a generalized form of tree where a parent term can have multiple child terms, and also a child term can have multiple parent terms. The DAG is directed and a link connects from a child term to a parent term, representing “the child is a sub-class of the parent.” (left panel). In some other cases, the direction can also be reversed to represent “a parent includes the child” (right panel).
In this vignette, I will introduce the ontology_DAG class as well as related functions with a tiny example. As shown in the following diagram, there are six terms in the DAG where term “a” is the root, term “e” and “f” are two leaf terms. Note term “c” has two parents.
Construct the object
The DAG object is constructed via a list of parent-child pairs. The following code constructs a DAG in the diagram above.
library(simona)
parents = c("a", "a", "b", "b", "c", "d")
children = c("b", "c", "c", "d", "e", "f")
dag = create_ontology_DAG(parents, children)Typing dag prints the basic information of the DAG.
dag## An ontology_DAG object:
## Source: Ontology
## 6 terms / 6 relations
## Root: a
## Terms: a, b, c, d, ...
## Max depth: 3
## Avg number of parents: 1.20
## Avg number of children: 1.00
## Aspect ratio: 0.67:1 (based on the longest distance from root)
## 0.68:1 (based on the shortest distance from root)Aspect ratio is calculated as width/height, where width is the largest number of terms on a specific depth (i.e. max(table(depth(dag)))). Definition of the height of a term in the DAG depends on whether using the longest or the shortest distance from root. The aspect ratio gives an impression of how the shape of the DAG looks like (fat or slim).
The ontology_DAG object can also be constructed by a vector of parent-child links:
dag = create_ontology_DAG(c("a-b", "a-c", "b-c", "b-d", "c-e", "d-f"))Following functions return the root term, leaf terms and test whether terms are leaves.
dag_root(dag)## [1] "a"
dag_leaves(dag)## [1] "e" "f"
dag_is_leaf(dag, letters[1:6])## a b c d e f
## FALSE FALSE FALSE FALSE TRUE TRUEdag_all_terms() returns a vector of all terms. dag_n_terms() simply returns the number of all terms in the DAG.
dag_all_terms(dag)## [1] "a" "b" "c" "d" "e" "f"
dag_n_terms(dag)## [1] 6DAG traverse
Numbers of child/parent/offspring/ancestor terms.
n_children(dag)## a b c d e f
## 2 2 1 1 0 0
n_parents(dag)## a b c d e f
## 0 1 2 1 1 1
n_offspring(dag)## a b c d e f
## 5 4 1 1 0 0
n_ancestors(dag)## a b c d e f
## 0 1 2 2 3 3n_connected_leaves() returns numbers of leaves that every term can reach (or has a finite directed distance to). Leaf terms have value of 1.
n_connected_leaves(dag)## a b c d e f
## 2 2 1 1 1 1Parents/children of a single term or union of parents/children of a group of terms. The term argument can be a single term name or a vector of term names.
dag_parents(dag, "c")## [1] "a" "b"
dag_parents(dag, c("d", "e")) # union of d's parents and e's parents## [1] "b" "c"
dag_children(dag, "b")## [1] "c" "d"It is similar to get offspring/ancestor terms of a single term or a group of terms.
dag_offspring(dag, "b")## [1] "c" "d" "e" "f"
dag_ancestors(dag, "e")## [1] "a" "b" "c"In many methods, when constructing the set of a term’s ancestors of offsprings, the set itself is also included. In dag_offspring() and dag_ancestors(), an argument include_self can be set to TRUE to include the term itself.
dag_offspring(dag, "b", include_self = TRUE)## [1] "b" "c" "d" "e" "f"
dag_ancestors(dag, "e", include_self = TRUE)## [1] "a" "b" "c" "e"Reorder
DAG can be reordered by adjusting the order of child terms under every term. There are two ways to reorder the DAG. 1. Reorder the DAG by a vector of numeric values that correspond to every term in the DAG; 2. Reorder by a vector of numeric values that correspond to every leaf term in the DAG.
The two ways are very similar. If the vector corresponding to the complete set of terms is provided, each term in the DAG is associated to a score which is the mean value of all its offspring terms (including itself) in the DAG; while if the a vector of leaf-level value is provided, each term in the DAG is associated to a score which is the mean of all its connectable leaf terms in the DAG.
Taking dag as an example, term c is on the left of term d
dag_children(dag, "b")## [1] "c" "d"First, a numeric vector for all terms in the DAG is provides to reorder dag. Note that the order of value should correspond to terms in dag_all_terms(dag).
dag2 = dag_reorder(dag, value = c(1, 1, 10, 1, 10, 1))
dag_children(dag2, "b")## [1] "d" "c"Or a numeric vector for all leaf terms. Note that the order of value here should correspond to terms in dag_leaves(dag).
dag3 = dag_reorder(dag, value = c(10, 1))
dag_children(dag3, "b")## [1] "d" "c"dag_permutate_children() randomly permutes child terms under every term.
dag2 = create_ontology_DAG(c("a-b", "a-c", "a-d", "a-e", "a-f"))
dag3 = dag_permutate_children(dag2)
dag_children(dag3, "a")## [1] "b" "e" "c" "f" "d"
dag3 = dag_permutate_children(dag2)
dag_children(dag3, "a")## [1] "d" "c" "b" "e" "f"Distance
The depth of a term is defined as the longest distance from root and is calculated by dag_depth(). If you want the shortest distance from root, use dag_shortest_dist_from_root().
dag_depth(dag)## a b c d e f
## 0 1 2 2 3 3## a b c d e f
## 0 1 1 2 2 3Similarly, the “height” of a term is defined as the longest distance to its reachable leaf terms. dag_shortest_dist_to_leaves() returns the shortest distance to leaves.
dag_height(dag)## a b c d e f
## 3 2 1 1 0 0## a b c d e f
## 2 2 1 1 0 0These four functions introduced so far calculate distance from root or to leaves for all terms in the DAG. The following functions are more general which calculates distance from a self-defined group of terms or until a group of terms.
dag_depth() is identical to dag_longest_dist_to_offspring(dag, root), and dag_height() is identical to dag_longest_dist_from_ancestors(dag, leaves) where root is a single root term and leaves is a vector of all leaf terms in the DAG.
dag_longest_dist_to_offspring(dag, from)
dag_shortest_dist_to_offspring(dag, from)
dag_longest_dist_from_ancestors(dag, to)
dag_shortest_dist_from_ancestors(dag, to)Please note, these four functions are applied to all terms. They return a vector with the same length as the total terms in the DAG. If a term is not included in the calculation, e.g. the root term when from does not include it, dag_longest_dist_to_offspring() and dag_shortest_dist_to_offspring() will assign -1 to the root term.
Given any two terms in the DAG, the following four functions calculate their pair-wise distance. There are different ways to calculate the distance:
For two terms \(a\) and \(b\), shortest_distances_via_NCA() calculates the distance as:
\[ \min_{t \in \mathrm{CA}(a, b)}(D_{sp}(t, a) + D_{sp}(t, b)) \]
where \(\mathrm{CA}(a, b)\) is the set of common ancestors (CA) of \(a\) and \(b\). \(D_{sp}(x, y)\) is the shortest distance between \(x\) and \(y\). In this way, common ancestor \(t\) which returns the minimal distance between \(a\) and \(b\) is called the “nearest common ancestor (NCA)” of \(a\) and \(b\). The usage of the function is as follows. It returns a symmetric matrix of pair-wise distance of terms.
shortest_distances_via_NCA(dag, terms)longest_distances_via_LCA() calculates the distance as:
\[ \mathrm{len}(t, a) + \mathrm{len}(t, b) \]
where \(\mathrm{len}(x, y)\) is the longest distance between \(x\) and \(y\). Common ancestor \(t\) is the one with the largest depth in DAG and it is called the lowest common ancestor (LCA) of \(a\) and \(b\):
\[ t = \operatorname*{argmax}_{t \in \mathrm{CA}(a, b)} \delta(t)\]
where \(\delta(t)\) is the depth (maximal distance from root) of term \(t\). The usage of the function is:
longest_distances_via_LCA(dag, terms)The next two functions treat the DAG relations as directional. There is a positive distance value only if a term is an ancestor of the other, or else the distance is set to -1 in the returned distance matrix.
shortest_distances_directed(dag, terms)
longest_distances_directed(dag, terms)Convert to other formats
To an igraph object
DAG is a graph. dag_as_igraph() converts the DAG to an igraph object.
g = dag_as_igraph(dag)
g## IGRAPH 1fa88b9 DN-- 6 6 --
## + attr: name (v/c)
## + edges from 1fa88b9 (vertex names):
## [1] a->b a->c b->c b->d c->e d->fDraw the graph with a hierarchical graph layout:
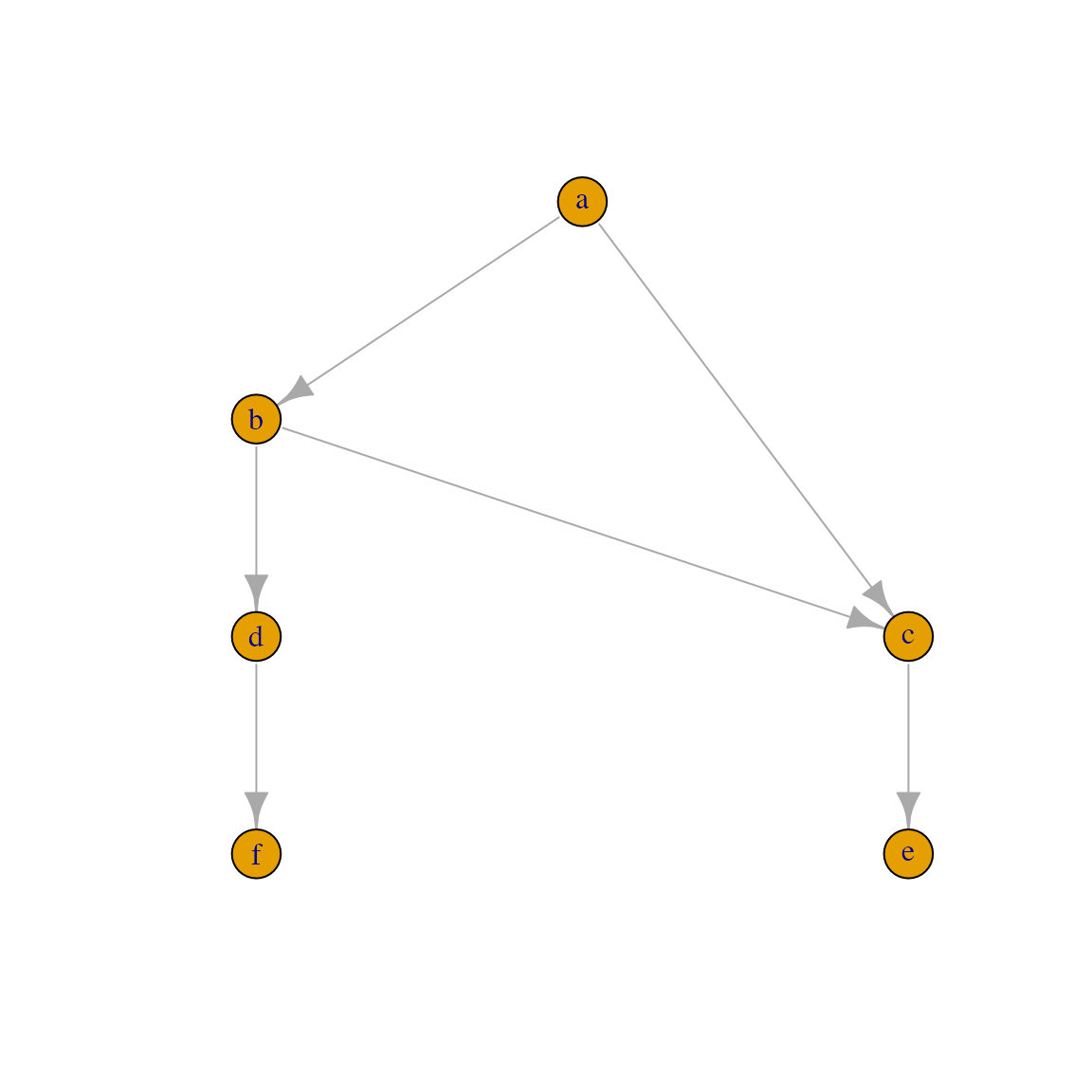
To a dendrogram object
In a DAG, a term can have multiple parents. A tree is a reduced form of a DAG where a term only has one parent. dag_treelize() simplifies a DAG to a tree. The reducing is applied in a breadth-first manner:
Starting from root and on a certain depth, for every term \(a\) on this depth, its child term \(c\) and parent-child relation are kept only when \(\delta_c = \delta_a + 1\). If \(c\) is selected, it is marked as visited and will not be checked again. In this way, depths of all terms in the orignal DAG are still identical to the depths in the tree.
tree = dag_treelize(dag)
dag_depth(dag)## a b c d e f
## 0 1 2 2 3 3
dag_depth(tree)## a b c d e f
## 0 1 2 2 3 3Note, although we are talking about “a tree”, tree in the example is still an ontology_DAG object where only multi-parent relations are removed. If n_relations + 1 == n_terms, there is special mark of being a tree in the object.
tree## An ontology_DAG object:
## Source: Ontology, treelized
## 6 terms / 5 relations / a tree
## Root: a
## Terms: a, b, c, d, ...
## Max depth: 3
## Aspect ratio: 0.67:1When the DAG is a tree, it can be converted to a dendrogram object. Since some nodes may only have one child, I add labels on nodes to mark the nodes are there. You can see the link a->c is removed from the DAG.
dend = dag_as_dendrogram(tree)
dend = dendrapply(dend, function(d) {
attr(d, "nodePar") = list(pch = attr(d, "label"))
d
})
plot(dend, leaflab = "none")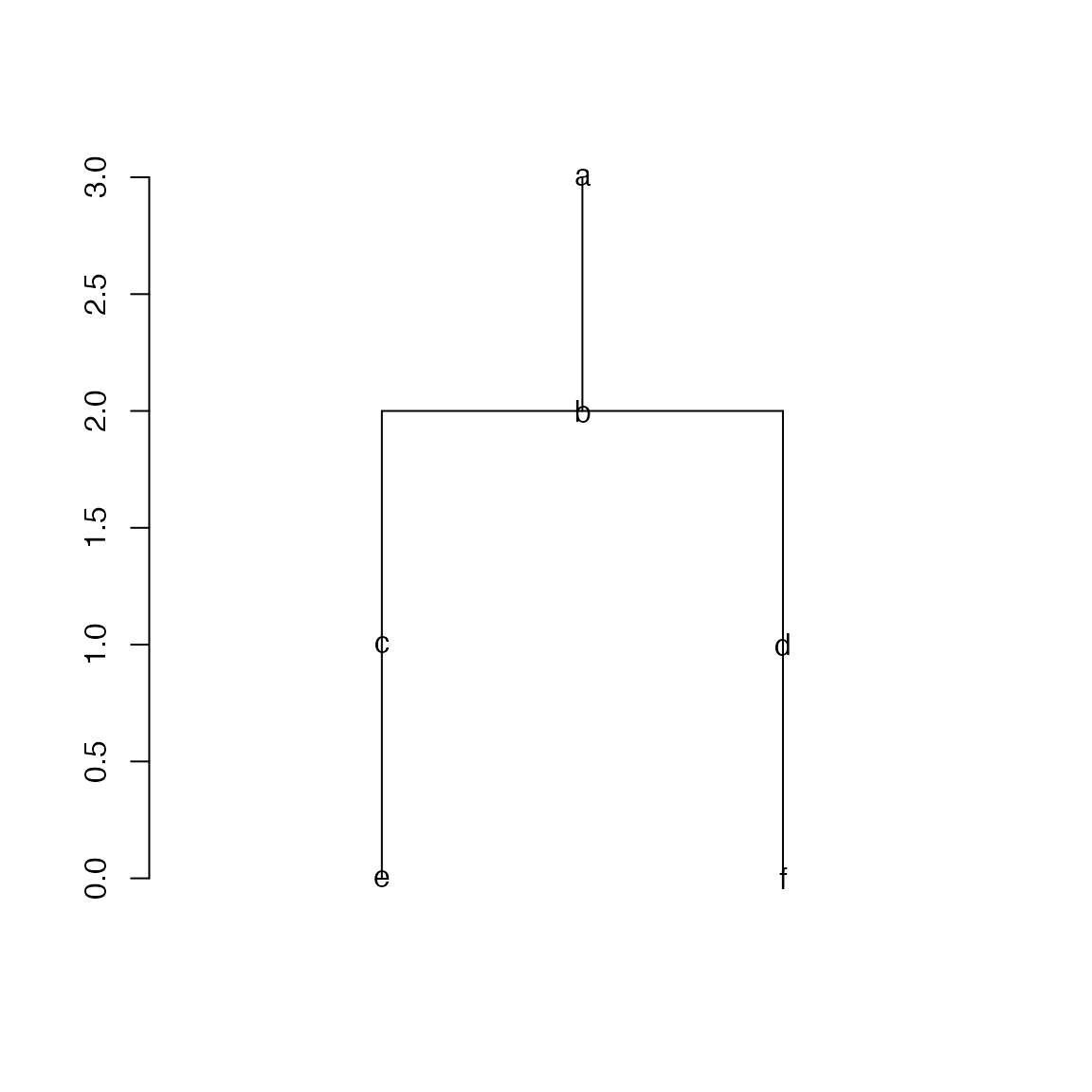
Note reducing DAG is mainly for visualization purpose.
Add relation types
If relations argument is not set, all relations are treated as in “is_a” such that a child is a sub-class of a parent class. It is common in ontologies, besides the “is_a” relation, there are many other self-defined relation types. One typical relation type is “part_of”. A vector of relation types can be set via the relations argument. It might be used later when calculating semantic similarities where different relation types are assigned with different weights.
relations = c("is_a", "is_a", "part_of", "part_of", "is_a", "is_a")
dag = create_ontology_DAG(parents, children, relations = relations)
dag## An ontology_DAG object:
## Source: Ontology
## 6 terms / 6 relations
## Root: a
## Terms: a, b, c, d, ...
## Max depth: 3
## Avg number of parents: 1.20
## Avg number of children: 1.00
## Aspect ratio: 0.67:1 (based on the longest distance from root)
## 0.68:1 (based on the shortest distance from root)
## Relations: is_a, part_ofAdd annotations
Terms can have external items annotated. One typical example is that GO terms can have genes annotated. Later the annotated items can be used for calculating semantic similarities.
The annotation should be set as a list of character vectors, where names of the element vectors should correspond to term names in the DAG so that annotations can be mapped to terms.
annotation = list(
"a" = c("t1", "t2", "t3"),
"b" = c("t3", "t4"),
"c" = c("t5"),
"d" = c("t7"),
"e" = c("t4", "t5", "t6", "t7"),
"f" = c("t8")
)
dag = create_ontology_DAG(parents, children, annotation = annotation)
dag## An ontology_DAG object:
## Source: Ontology
## 6 terms / 6 relations
## Root: a
## Terms: a, b, c, d, ...
## Max depth: 3
## Avg number of parents: 1.20
## Avg number of children: 1.00
## Aspect ratio: 0.67:1 (based on the longest distance from root)
## 0.68:1 (based on the shortest distance from root)
## Annotations: 8 items
## t1, t2, t3, t4, ...Due to the nature of DAG, if a child term is annotated to an item, all its ancestor terms are also associated with that item. The calculation of annotated items is applied in a recursive way.
For a term \(x\), denote \(\mathcal{C}_x\) is the set of its child terms, the items “directly” annotated to \(x\) denoted as set \(G^*_x\), then the final set of items annotated to \(x\) denoted as \(G_x\) is the union of all items annotated to its child terms as well as \(x\) itself.
\[ G_x = \left( \bigcup_{z \in \mathcal{C}_x} G_z \right) \bigcup G^*_x \]
Note above equation is applied resursively. If denoting \(\mathcal{D}^+_x\) as the set of \(x\)’s offspring terms plus \(x\) itself, \(G^*_x\) can also be written as:
\[ G_x = \bigcup_{z \in \mathcal{D}^+_x} G_z^* \]
The numbers of annotated items of DAG terms can be obtained via the function n_annotations(). The attribute attr(,"N") is the maximal number of items annotated to the DAG (the same as max(n_annotations(dag))), which normally corresponds to the root term.
n_annotations(dag)## a b c d e f
## 8 6 4 2 4 1
## attr(,"N")
## [1] 8The next two functions return the associations between terms and items.
term_annotations(dag, letters[1:6])## $a
## [1] "t1" "t2" "t3" "t4" "t5" "t7" "t6" "t8"
##
## $b
## [1] "t3" "t4" "t5" "t7" "t6" "t8"
##
## $c
## [1] "t4" "t5" "t7" "t6"
##
## $d
## [1] "t7" "t8"
##
## $e
## [1] "t4" "t5" "t7" "t6"
##
## $f
## [1] "t8"
annotated_terms(dag, c("t1", "t2", "t3"))## $t1
## [1] "a"
##
## $t2
## [1] "a"
##
## $t3
## [1] "a" "b"Or return a binary matrix:
term_annotations(dag, letters[1:6], return = "matrix")## t1 t2 t3 t4 t5 t7 t6 t8
## a 1 1 1 1 1 1 1 1
## b 0 0 1 1 1 1 1 1
## c 0 0 0 1 1 1 1 0
## d 0 0 0 0 0 1 0 1
## e 0 0 0 1 1 1 1 0
## f 0 0 0 0 0 0 0 1
annotated_terms(dag, c("t1", "t2", "t3"), return = "matrix")## a b c d e f
## t1 1 0 0 0 0 0
## t2 1 0 0 0 0 0
## t3 1 1 0 0 0 0Pseudo root
The DAG should be lead by a single root term. If in an ontology there are multiple root terms, a pseudo root named "~all~" is automatically added. You can see this process from the message of the function call.
parents = c("a", "a", "b", "x", "x", "y")
children = c("b", "c", "c", "z", "y", "z")
create_ontology_DAG(parents, children)## There are more than one root:
## a, x
## A super root (~~all~~) is added.## An ontology_DAG object:
## Source: Ontology
## 7 terms / 8 relations
## Root: ~~all~~
## Terms: a, b, c, x, ...
## Max depth: 3
## Avg number of parents: 1.33
## Avg number of children: 0.80
## Aspect ratio: 0.67:1 (based on the longest distance from root)
## 2:1 (based on the shortest distance from root)Alternative terms
During the development of an ontology, old terms may be replaced by new terms. In the .obo format, there are the following three tags: alt_id, replaced_by and consider. The last two tags are used especially for obsolete (removed) terms. If there is only one-to-one mapping between the alternative term IDs and DAG term IDs, the mapping can be provided as a named vector. The mapping variable is filtered and saved in the alternative_terms slot of the DAG object.
parents = c("a", "a", "b", "b", "c", "d")
children = c("b", "c", "c", "d", "e", "f")
alt = c("old_a" = "a", "old_b" = "b")
dag = create_ontology_DAG(parents, children, alternative_terms = alt)
dag@alternative_terms## old_a old_b
## "a" "b"If the mapping is one-to-many, the mapping variable should be provided as a named list. There are several filterings. First, the non-DAG term IDs in the mapping variable are removed. If the mapping for an alternative ID is still one-to-many, the first DAG term is used.
alt = list("old_a" = c("a", "b", "x"), "old_y" = "y")
dag = create_ontology_DAG(parents, children, alternative_terms = alt)
dag@alternative_terms## old_a
## "a"Sub-DAG
The following code returns a sub-DAG where the input term is picked as the root of the sub-DAG.
# or with the double bracket: dag[["b"]]
dag["b"]## An ontology_DAG object:
## Source: Ontology
## 5 terms / 4 relations / a tree
## Root: b
## Terms: b, c, d, e, ...
## Max depth: 2
## Aspect ratio: 1:1Two indicies can be provided in the brackets where the first one corresponds to root terms and the second one corresponds to leaf terms:
# the same as dag["b"], a sub-DAG where b is the root dag["b", ] # a sub-DAG where b is the root and e is the only leaf dag["b", "e"] # a sub-DAG that contains all e's ancestors and e itself dag[, "e"]
With the more general function dag_filter(), you can obtain a sub-DAG by providing a group of terms, a subset of relations, a group of root terms or a leaf terms. Note if there are multiple root after the fitlering, a pseudo node will be automatically added.
dag_filter(dag, terms, relations, root, leaves)
Meta data frame
The DAG object can have a meta data frame attached, which provides more information on the terms in the DAG. Examples can be found from the “2. Gene ontology” vignette. dag_filter() can also be applied based on columns in the meta data frame.
Other aspects
When creating the ontology_DAG object, create_ontology_DAG() checks whether there exist cyclic links. Denote a cyclic link as a->b->c->d->b. The last link d->b is automatically removed to get rid of cyclic links.
A second, even more extreme scenario are isolated sub-graphs represented as rings where the root can not be identified there. For example a->b->c->d->a is a ring where there is no root term, thus, it cannot be attached to the main DAG by adding a pseudo root. Rings with the form of a->b->a is more observed in public ontology datasets. create_ontology_DAG() automatically removes rings.
Session info
## R version 4.3.3 (2024-02-29)
## Platform: x86_64-apple-darwin20 (64-bit)
## Running under: macOS Sonoma 14.6.1
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRlapack.dylib; LAPACK version 3.11.0
##
## locale:
## [1] en_GB.UTF-8/en_GB.UTF-8/en_GB.UTF-8/C/en_GB.UTF-8/en_GB.UTF-8
##
## time zone: Europe/Berlin
## tzcode source: internal
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] igraph_2.0.3 simona_1.3.12 knitr_1.45
##
## loaded via a namespace (and not attached):
## [1] sass_0.4.9 xml2_1.3.6 shape_1.4.6.1
## [4] digest_0.6.35 magrittr_2.0.3 evaluate_0.23
## [7] grid_4.3.3 RColorBrewer_1.1-3 iterators_1.0.14
## [10] circlize_0.4.16 fastmap_1.1.1 foreach_1.5.2
## [13] doParallel_1.0.17 jsonlite_1.8.8 GlobalOptions_0.1.2
## [16] promises_1.3.0 ComplexHeatmap_2.18.0 purrr_1.0.2
## [19] codetools_0.2-19 textshaping_0.3.7 jquerylib_0.1.4
## [22] cli_3.6.2 shiny_1.8.1.1 rlang_1.1.3
## [25] crayon_1.5.2 scatterplot3d_0.3-44 cachem_1.0.8
## [28] yaml_2.3.8 tools_4.3.3 parallel_4.3.3
## [31] memoise_2.0.1 colorspace_2.1-0 httpuv_1.6.15
## [34] GetoptLong_1.0.5 BiocGenerics_0.48.1 mime_0.12
## [37] vctrs_0.6.5 R6_2.5.1 png_0.1-8
## [40] matrixStats_1.3.0 stats4_4.3.3 lifecycle_1.0.4
## [43] S4Vectors_0.40.2 fs_1.6.4 htmlwidgets_1.6.4
## [46] IRanges_2.36.0 clue_0.3-65 ragg_1.3.1
## [49] cluster_2.1.6 pkgconfig_2.0.3 desc_1.4.3
## [52] later_1.3.2 pkgdown_2.0.9 bslib_0.7.0
## [55] glue_1.7.0 Rcpp_1.0.12 systemfonts_1.0.6
## [58] highr_0.10 xfun_0.43 xtable_1.8-4
## [61] rjson_0.2.21 htmltools_0.5.8.1 rmarkdown_2.26
## [64] Polychrome_1.5.1 compiler_4.3.3